Challenges in realising the potential of wastewater-based epidemiology to quantitatively monitor and predict the spread of disease
Abstract
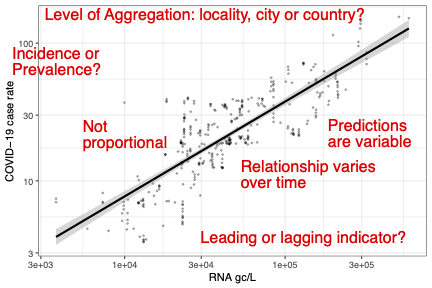
Researchers around the world have demonstrated correlations between measurements of SARS-CoV-2 RNA in wastewater (WW) and case rates of COVID-19 derived from direct testing of individuals. This has raised concerns that wastewater-based epidemiology (WBE) methods might be used to quantify the spread of this and other diseases, perhaps faster than direct testing, and with less expense and intrusion. We illustrate, using data from Scotland and the USA, the issues regarding the construction of effective predictive models for disease case rates. We discuss the effects of variation in, and the problem of aligning, public health (PH) reporting and WW measurements. We investigate time-varying effects in PH-reported case rates and their relationship to WW measurements. We show the lack of proportionality of WW measurements to case rates with associated spatial heterogeneity. We illustrate how the precision of predictions is affected by the level of aggregation chosen. We determine whether PH or WW measurements are the leading indicators of disease and how they may be used in conjunction to produce predictive models. The prospects of using WW-based predictive models with or without ongoing PH data are discussed.